AsianScientist (Jul. 30, 2019) – By probing the mechanisms that allow the bacteria Pseudomonas aeruginosa to regulate gene expression and virulence, researchers at the City University of Hong Kong (CityU), China, hope to discover new compounds for treating drug-resistant microbes. Their findings are published in the journal Nature Communications.
The superbug P. aeruginosa is increasingly common in hospitals across the globe, and immunocompromised patients are particularly susceptible to infection. The pathogen is also naturally tolerant to many clinically important antibiotics such as ampicillin, amoxicillin and vancomycin. In 2017, the World Health Organization classified this notorious bacterium as one of the three ‘critical priority pathogens’ for which new drugs are urgently needed.
In the present study, scientists led by Assistant Professor Deng Xin and Associate Professor Wang Xin at City U uncovered the genomic regulatory network in P. aeruginosa and identified the master regulators determining pathogenicity.
They found that in P. aeruginosa, numerous transcription factors—proteins that regulate the expression of other genes—regulate virulence by tuning quorum sensing, a molecular mechanism that bacteria use to communicate with one another. Their findings also implicated the type III and type VI secretion system of P. aeruginosa, consisting of microscopic structures that the microbes use to promote infection.
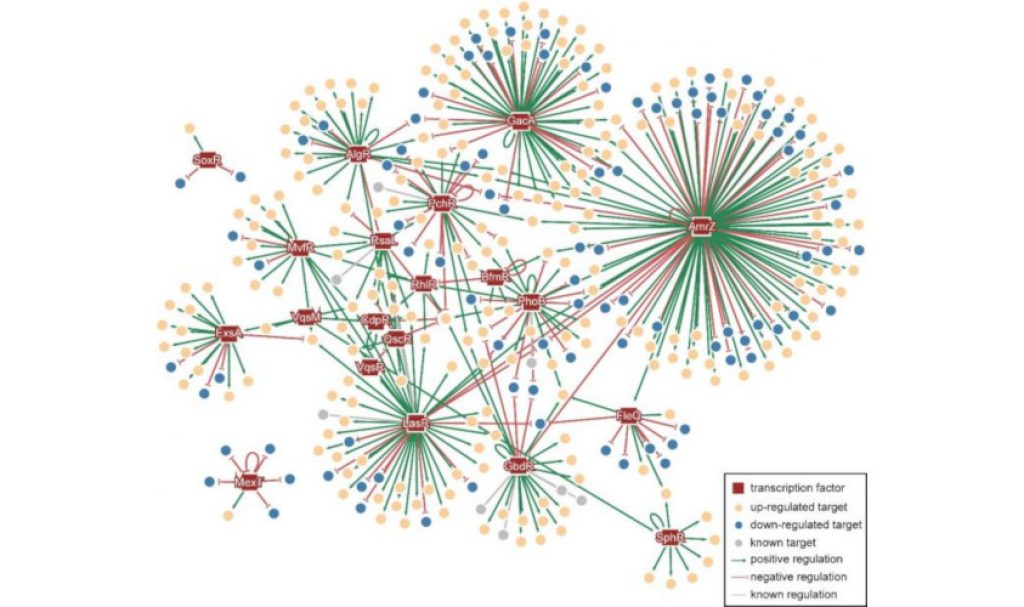
To obtain a global perspective on possible pathogenic pathways and discover new potential drug targets in Pseudomonas aeruginosa, the team next created the P. aeruginosa Genomic regulatory network, or PAGnet. In essence, PAGnet is a map of the regulatory relationships among 20 transcription factors and their functional target genes, useful for providing researchers with an overview of possible mechanisms of virulence in P. aeruginosa.
“The master regulators we identified are potential antibiotic targets, which has important clinical significance for the development of new antibiotics for treating P. aeruginosa in the future. More importantly, the network we built is not just for P. aeruginosa; the methodology and conclusions of this work may be applicable to other bacterial pathogens as well,” said Deng.
The article can be found at: Huang et al. (2019) An Integrated Genomic Regulatory Network of Virulence-related Transcriptional Factors in Pseudomonas aeruginosa.
———
Source: City University of Hong Kong.
Disclaimer: This article does not necessarily reflect the views of AsianScientist or its staff.