A hundred million genomes
But just how much more data is required? When pressed for an estimate, Wang says that iCarbonX’s goal is to sequence the genomes of 100 million people.
“Not only that, we want to have multi-omics data as well, and the data collection has to be at multiple points over each individual’s life span; it has to be longitudinal data. There are, of course, many difficulties in collecting such multi-layered ‘omics data for millions of individuals, but this is something you just have to do,” Wang says.
“I cannot tell you exactly how many ‘omics we will be tracking, but I can tell you that iCarbonX is currently doing a lot of offline set-up and quality control aimed at a consumer pipeline for sample collection. We are also evaluating several proteomics and metabolomics technologies that are not as mature as DNA sequencing,” he shares.
To achieve this ambitious aim, iCarbonX already has US$106 million at its disposal from a Series A round of funding in April 2016. As a “first step,” Wang estimates that the company will need about US$2 billion in funding. When BGI participated in the Human Genome Project, the cost of sequencing a single genome was US$3 billion (and took 13 years!). In 2014, Illumina broke the psychological US$1,000 barrier with its HiSeq X Ten machine which promised to sequence a complete genome for that price or slightly less.
“The reagent cost for sequencing is now about US$200 [per genome]. I think that it will soon reach US$50 in the next couple of years, and then go down to US$10. Eventually, it will be free, if not in five years then a little more than five years,” Wang predicts.
“Come to us; we will sequence you for free! If the genomic data is free, the epigenome, metabolome and proteome should all be free, too. Data collection will be free, but of course if you want a diagnostic report, a product or a drug, you will have to pay for it.
“The iCarbonX business model is essentially [that of] a data company; we will operate the data that feeds back into personalized services and products for consumers,” he explains.
From observation to prediction
The point of collecting all this data, Wang says, is to ultimately have enough predictive power to conduct preventive healthcare. He envisions a future where machine learning will be able to create digital versions of each person; an avatar where different life-extending interventions can be tested.
“The idea is to develop a ‘crystal ball’ or a ‘healthcare GPS’ that will serve as a guide to making the right lifestyle choices based on your personal genome and all the ‘omics and other data,” Wang continues.
“In such a scenario, precision nutrition will become essential. The advice will not just be ‘eat more vegetables’ or ‘eat less red meat,’ but it will also tell you specifically which vegetables you should eat and in what quantities.”
Indeed, Wang feels that this marriage of artificial intelligence and biology is a natural progression for the field as it matures. Noting how biology began as an observational discipline concerned with simply documenting and classifying the natural world, he points to the molecular biology revolution as a key inflexion point where biology began to be heavily experimental and hypothesis-driven.
“But by the time I founded BGI, [biology] had become more theoretical. People were starting to be data-driven, sequencing tens of thousands of individuals without any hypothesis in mind and from there, trying to understand the human genome,” Wang says.
As a proof-of-concept study to demonstrate the predictive power of multi-layered ‘omics data, Wang and his team at BGI collected over 3,000 different strains of foxtail millet (Setaria italica), growing them in the same field and collecting a range of ‘omics data.
“The results were really breathtaking,” Wang enthuses. “Our model allowed us to predict the phenotype of any given strain of foxtail millet with over 95 percent accuracy. Although the foxtail millet genome is much smaller and simpler than the human genome, our study nevertheless shows that machine learning technology can give you a much clearer idea of what the phenotype will be.”
Wang is confident that predictive models for humans based on ‘omics data is not far off, and “within the next five to ten years.”
“When I first started out in artificial intelligence twenty years ago, the question I always asked myself was: ‘Can machines think?’ That is no longer in doubt, but has raised a second question, which is actually deeper: ‘Can computers think like human beings?’
“It’s a much harder question because we don’t really understand how human beings think. But I believe we will get there soon.”
This article was first published in the print version of Asian Scientist Magazine, July 2016.
———
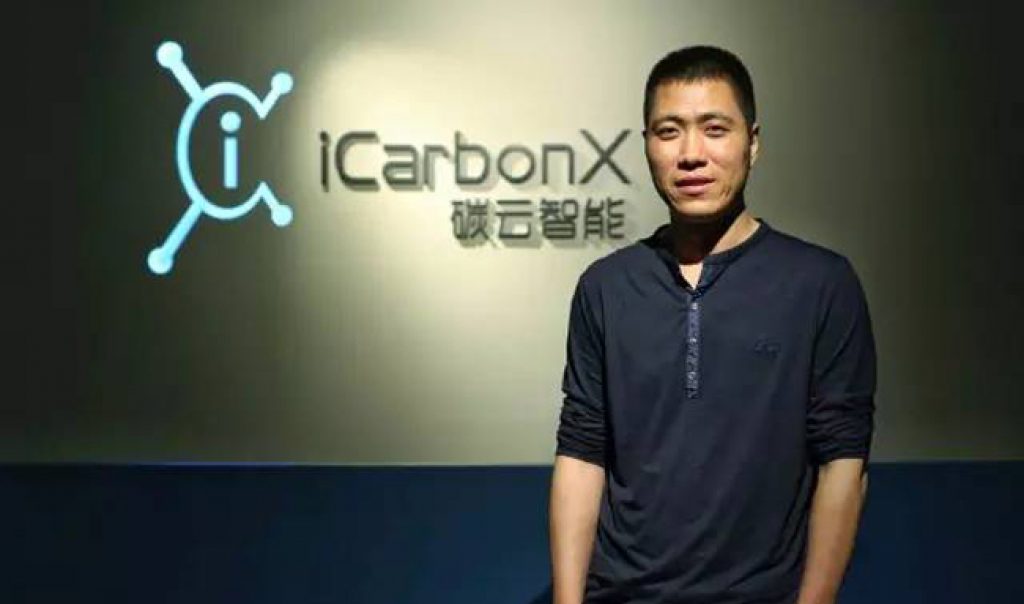
Copyright: Asian Scientist Magazine. Photo: iCarbonX.
To read more, subscribe to Asian Scientist Magazine in print and receive four issues of Asian Scientist Magazine delivered directly to your mailing address for 12 months, inclusive of taxes and postage.