AsianScientist (Feb. 17, 2016) – Although CRISPR-Cas9 technology has the potential to revolutionize molecular biology, it remains an imperfect tool, sometimes producing mutations known as indels at unintended (off-target) sites.
Now, researchers have developed a way of simultaneously mapping out the genome-wide specificities of multiple CRISPR-Cas9 nucleases to find both intentional and unwanted indels quickly and cheaply. Their method, called multiplex Digenome-seq (digested genome sequencing), has been described in Genome Research.
The team led by Professor Kim Jin-Soo from the Institute for Basic Science (IBS) had previously used CRISPR-Cas9 to edit the genome of plants without introducing foreign DNA. However, before CRISPR technology can be safely used in humans, scientists need to know the accuracy of the technique.
To profile the specificities of CRISPR-Cas9 nucleases, Kim and colleagues first digested human genomic DNA with Cas9 and multiple single guide RNA (sgRNA) sequences in vitro and then performed whole-genome sequencing.
“Digenome-seq differs from other cell-based methods in that it detects DNA cleavages in vitro, rather than in cells, using cell-free genomic DNA,” explained study first author Dr. Kim Daesik.
An advantage of using cell-free DNA is that it produces precise analysis results because chromatin, which is normally found in the cell, doesn’t interfere with the process. Also, because the DNA in question doesn’t need to be amplified en masse using PCR before investigation; Digenome-seq is easier, faster and cheaper to use than previous methods.
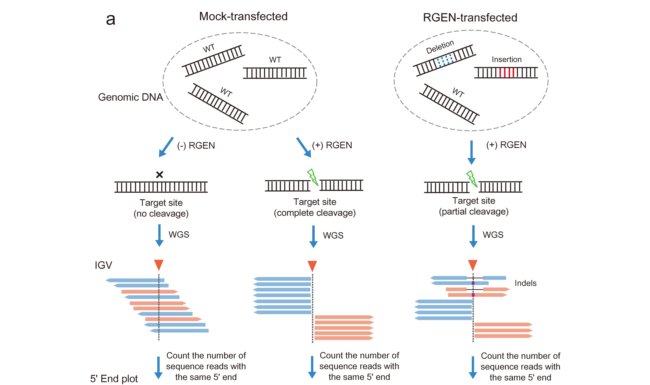
The sgRNA leads the Cas9 enzyme to both target and off-target sites on the DNA where it gets cleaved. By analyzing the cleaved sequences, the scientists found that sgRNAs transcribed from an oligonucleotide duplex caused many false-positive off-target sites. Multiplex Digenome-seq also captured many bona fide off-target sites that were missed by other genome-wide methods, at which indels were induced at frequencies <0.1 percent.
Unlike previously used processes that can only work with one sgRNA at a time, multiplex Digenome-seq performs the analysis of numerous sgRNAs to detect all of their on-target and off-target sites simultaneously. This multiplex process is faster, more efficient and more cost effective than previous methods.
“CRISPR-Cas9 is an incredible scientific breakthrough and will continue to be the research focus of biologists around the world,” said Kim, who is also the director of IBS. “This technique can be used to choose target sites with the least off-target effects, a method essential for therapeutic applications of CRISPR-Cas9.”
The article can be found at: Kim et al. (2016) Genome-wide Target Specificities of CRISPR-Cas9 Nucleases Revealed by Multiplex Digenome-seq.
———
Source: Institute for Basic Science; Photo: Shutterstock.
Disclaimer: This article does not necessarily reflect the views of AsianScientist or its staff.