AsianScientist (Oct. 7, 2020) – The impact of CRISPR gene editing has been nothing short of revolutionary. In less than a decade, it has spread across labs around the world and found its way into everything from growing better crops to eradicating faulty genes and even detecting viruses in a pandemic.
Given the scientific significance—and vast commercial potential—of CRISPR, it’s no surprise that the question of who really ‘discovered’ this game-changing technology has become the subject of a multimillion-dollar legal battle. Since 2012, the University of California, Berkeley (UC Berkeley) and the Broad Institute have been locked in a bitter feud over which institution owns the rights to CRISPR technology.
But when Professor Yoshizumi Ishino, then a freshly minted PhD graduate from Japan’s Osaka University, first came across a peculiar repetitive sequence in bacterial DNA in 1986, it was regarded as little more than an academic curiosity. In classic scientific understatement, the final line of the paper describing his findings reads “the biological significance of these sequences is not known.”

A strange sequence of events
If you were to take even the best-trained graduate student today and send them in a time machine back to the mid-1980s, they would find themselves at an utter loss in a lab like Ishino’s. Thermocyclers, those ubiquitous pieces of lab furniture taken for granted by modern scientists, were only invented in 1985. While getting a gene sequenced is now routine and easily done by an undergraduate assistant, back then sequencing a single gene was worthy of an entire PhD thesis.
The state-of-the-art gene sequencing method at the time was dideoxy nucleotide chain-termination or Sanger sequencing, a method that pieces the full sequence together from four separate reactions, one for each nucleotide: A, T, C or G.
“After each reaction, you had to run the sample on a gel, which took between two-and-a-half to six hours,” Ishino told Asian Scientist Magazine. “Then after electrophoresis, you dried the gel and exposed it to an X-ray film to get a radiograph to see the sequencing ladder. Finally, you had to read the result by hand and manually enter the sequence into a computer.”
Furthermore, due to limitations of the length of DNA that could be sequenced by this method, Ishino had to break up the relatively short gene he was studying—iap, encoding an isozyme-converting enzyme of Escherichia coli alkaline phosphatase—into smaller segments of a few hundred bases and stitch the full sequence together like a jigsaw puzzle from independent runs of the experiment.
While getting all this done using today’s sequencing technology would take little more than an afternoon, in 1986 it took Ishino months of working 14 hours a day.
“Because we used a radioactive label to identify the nucleotide sequences, I had to go to a special facility for radioisotopes. I was there from 8:30am till past 10pm every day, sometimes even staying there for several days in a row,” he said.
Despite the hard work required, the main project itself was straightforward enough.
“We completed the nucleotide sequencing, showed the gene products and how they worked on alkaline phosphatase. So the story was very clear and the paper was easily accepted,” Ishino said.
On hindsight, however, what was truly interesting about the study was what followed a few base pairs after: a stretch of 29 nucleotides repeated one after another five times in a row.
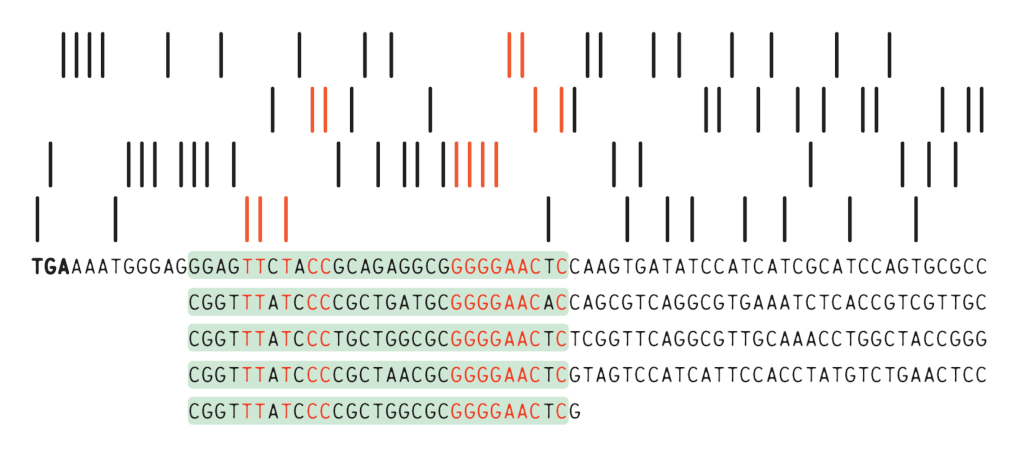
Like most scientists coming across an unexpected finding, Ishino at first thought that what he was seeing was a mistake. In the course of his experiments, Ishino found that many of the clones he had isolated contained a similar stretch of 29 nucleotides. Uncertain about how to interpret these findings—and further hampered by non-specific results that occurred because the dideoxy reaction would always stop when it encountered this specific sequence—Ishino repeated the experiment many times to be sure that what he was seeing was accurate enough to be published.
“After about half a year, I finally had the confidence to release my findings,” Ishino shared.
What was happening in the test tube was that the repetitive palindromic sequence at the end of iap was forming a stable stem-and-loop structure, preventing the dideoxy sequencing from completing correctly. When more sequences became available later by the E. coli genome project, this palindromic sequence was in fact repeated 14 times in total on the genome.
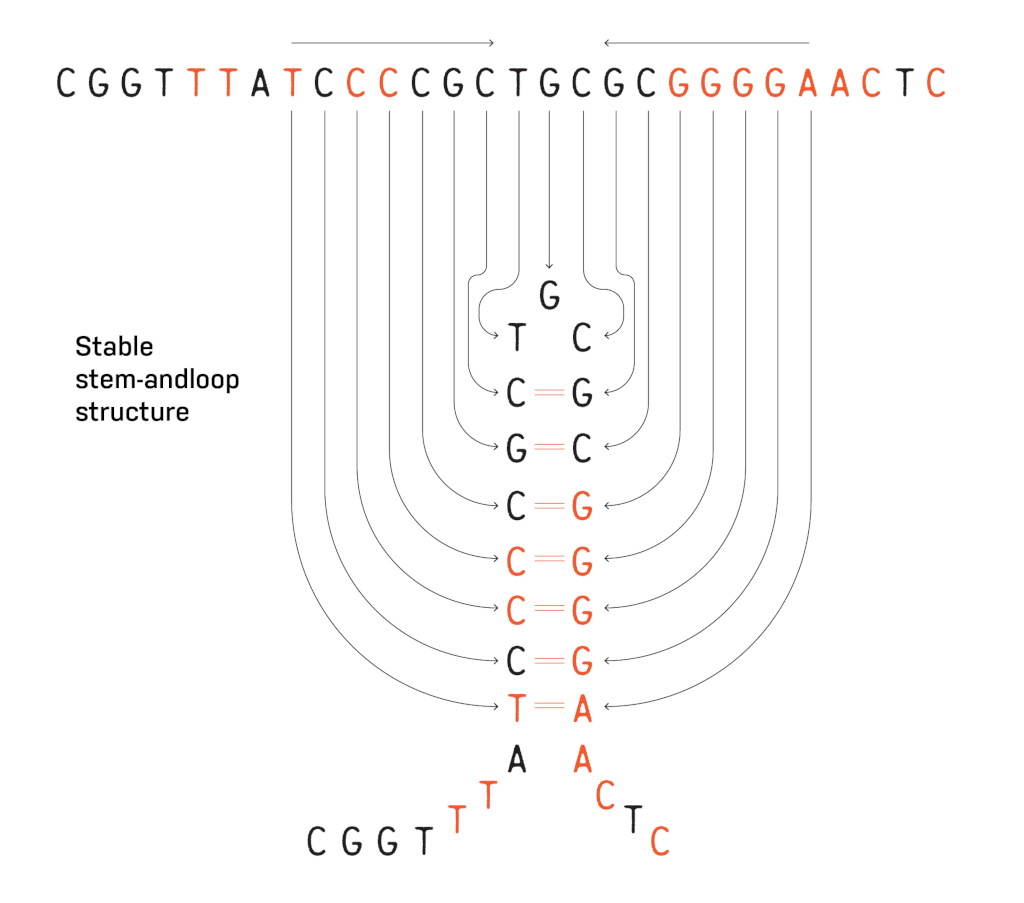
“The story about these strange repeats was not necessary to the paper, but they had never been found elsewhere before, so I added it in,” Ishino said. “It was exciting to me because the biological function was not yet known.”
Intrigued as he was about the purpose of these unusual repetitive sequences, Ishino had no means of taking the research further. The tools of molecular biology then were relatively limited, and besides, he had other plans for his future, including a post-doctoral position with Dieter Söll, a biochemist at Yale University famous for his work on transfer RNA.
“Because I had plans to go to Yale, I didn’t pursue it but said ‘another time,’” Ishino recounted.
Though he had no inkling of it at the time, the next opportunity would present itself over a decade later, when Ishino’s repeated palindromic sequences would finally get a name: clustered regularly interspaced short palindromic repeats or CRISPR.
Enchanted by (archaeal) enzymes
With experience at Yale under his belt, Ishino returned to Japan in 1989 to join the bioproducts development center at Takara Shuzo, a storied Japanese brewery that had spun out a successful biotechnology company producing enzymes. It was an exciting time to be in molecular biology, at the dawn of a new era enabled by methods such as polymerase chain reaction (PCR).
Like CRISPR decades later, PCR quickly established itself as an essential tool for molecular biology after being invented by American biochemist Kary Mullis in 1985. And well before Mullis was awarded the Nobel Prize for his ground-breaking invention in 1993, PCR was already the subject of legal challenges between the biotech giants of the day: Hoffman LaRoche, DuPont and Promega.
One key innovation at the heart of PCR—and at the center of most patent debates—is an enzyme called Taq polymerase. Before Taq polymerase was used, PCR required fresh polymerase enzymes to be added after each cycle of amplification, as the strand separation step required the reaction to be heated to over 95˚C, a temperature at which most enzymes would denature. Taq polymerase, which was isolated from an unusually heat-tolerant or thermophilic bacteria called Thermus aquaticus, changed the game by making PCR much simpler, cheaper and more efficient.
“I was in charge of developing PCR enzymes, which was what started my interest in hyperthermophiles, organisms that not only survive but thrive at extremely high temperatures,” Ishino said. “There were even organisms that could grow at over 100˚C; these were not bacteria, but archaea.”
Although they superficially resemble bacteria—and up till as recently as 1990 were in fact considered to be a kind of bacteria—Archaea are neither bacteria nor eukaryotes but represent the third fundamental domain of life on Earth. Possibly the most ancient form of life, archaea are single-celled organisms whose ancestors are thought to have been ingested by bacteria millions of years ago to eventually give rise to eukaryotes.
Fascinated by these under-studied organisms, Ishino mined Archaea for new enzymes to great effect, uncovering a constellation of interesting DNA polymerases and endonucleases.
“I discovered a new unique enzyme every few years; it was continuously exciting,” he said.
In a hyperthermophile archaeon called Pyrococcus furiosus, for instance, Ishino identified a new type of DNA polymerase that was only found in Archaea. Named PolD, it was later recognized as part of an entirely new family of DNA polymerases which could help us understand ancient DNA replication machinery. PolD also showed promise as a thermostable PCR enzyme that could potentially dethrone Taq polymerase.

By now, it was the mid-1990s and Ishino was beginning to establish himself as one of the leaders in Archaea research. Around the same time, in 1996, a landmark paper that would electrify the field was published by Carl Woese, the microbiologist who first called for Archaea to be recognized as a distinct domain, and Craig Venter, who would later go on to publish a draft sequence of the first human genome.
That paper, published in Science, described the first complete genome of an archaeon, conclusively showing that Archaea were distinct from both bacteria and eukaryotes. As Ishino read through the paper with great excitement, something striking immediately jumped out at him: 18 copies of a unique, repetitive sequence that reminded him of what he had seen in E. coli a decade before.
“They too couldn’t say anything about the function of these strange repeats, but noted that there were very many copies of that repeated sequence in the archaeon they had sequenced: Methanococcus jannaschii,” Ishino said.
It occurred to Ishino then that this could be his opportunity to return to those enigmatic palindromic sequences he had discovered at the start of his career.
“But at that point, I was so interested in the gene in M. jannaschii encoding the homolog of PolD that I originally discovered in Pyrococcus and trying to reconstitute an in vitro replication system that I forgot to come back to what later became known as CRISPR,” said Ishino, who by 2002 had been appointed a full professor at Kyushu University, where he still conducts research today.

And the beat goes on
While Ishino continued his research on Archaea, later branching out into metagenomics and astrobiology, others began taking note of the repeats he first observed in 1986. One of them was a young graduate student, Francisco Mojica, who also noticed the unusual sequences in 1992, naming them tandem repeats or TREPS on the suggestion of his advisor. Though the sequences now had a name, nobody had any clue what they were for, with Mojica incorrectly suggesting that they were involved in segregating DNA during cell division.
As science entered the genomics age with the increasing accessibility of genetic sequencing, more and more repeated structures began to be found across a wide range of microorganisms, with each research group giving them their own name. It was Ruud Jansen at Utrecht University, Netherlands—in correspondence with Mojica—who finally settled on clustered regularly interspaced short palindromic repeats, putting the term ‘CRISPR’ into the literature for the first time in 2002. As it turned out, Ruud was not so much interested in the CRISPR sequences themselves but the enzymes typically found shortly after, DNA-cutting proteins that he called CRISPR-associated proteins or Cas.
It was only in 2005, however, that scientists began to have an inkling about what CRISPR was doing in bacteria. That year, three independent groups—including Mojica’s at the University of Alicante, Spain—came to the realization that the spacer sequences between the CRISPR repeats resembled virus sequences.
The next piece of the puzzle fell into place when researchers at Danisco, the Copenhagen-based yoghurt maker, showed that the spacer sequences gave bacteria the ability to resist infections of the viruses that the sequences matched. In other words: CRISPR was actually a kind of bacterial immune defense system. Ishino, like other researchers interested in CRISPR, was gobsmacked.
“I never imagined that it would apply to the immune system,” he said.
But what scientists would later put this bacterial immune system to use for would be even more astounding, sparking a revolution in molecular biology.
The three revolutions in molecular biology
How exactly does the CRISPR-Cas system protect bacteria from invading viruses? Subsequent work, including from the laboratory of Jennifer Doudna at UC Berkeley, showed that bacteria use the spacer sequences as a reference, translating the DNA into RNA that is then taken up by Cas proteins. If this complex comes across a matching sequence of viral genetic material, the Cas protein cuts up the virus DNA, thereby preventing it from replicating in the host.
The key innovation, by Doudna and others including Zhang Feng and George Church at the Broad Institute, was to use this ability to cut DNA in other contexts, namely in vitro and in human cells. While cutting up the DNA of viruses simply destroys it, cutting DNA in a cellular context triggers repair mechanisms that can mend broken DNA. By exploiting these DNA repair mechanisms, researchers were in effect able to both cut and paste DNA at will. In short, CRISPR enabled scientists to re-write the code of life.
Zhang, Doudna and her collaborator Emmanuelle Charpentier at the Max Planck Institute for Infection Biology are now among the most famous and feted scientists in the world, with rumors swirling in their wake during Nobel Prize season every year. Though well respected in his own right, Ishino is lesser known outside Archaea circles. But looking back at his four decades of science, he has no regrets.
“If I could go back to 1987, maybe I should have studied more about CRISPR,” Ishino said. “If I had stayed in Osaka, I would definitely have continued that research; that was my feeling.”
“But when I look back on my research over 40 years, I feel happy to have contributed to three revolutions in genetic engineering technology,” he continued. “During the first revolution, I contributed to the research on restriction enzymes and DNA ligases, which have been used as the first genetic engineering technology to ‘cut and paste’ DNA. During the second revolution, I developed useful enzymes, which have been practically used for PCR. And now we are in the third revolution of genetic editing with CRISPR.”

Ishino continues to contribute to the current CRISPR revolution and has in fact found several new Cas proteins from his metagenomics studies. Drawing on his experience with developing PCR enzymes as alternatives to Taq polymerase, which required royalties to be paid to Hoffman LaRoche (later Roche), Ishino thinks alternatives to the most common Cas protein, Cas9, would turn the CRISPR-Cas system into something like what we have for restriction enzymes, where you can pick the enzyme of your choice based on what you need it to do.
“If you develop a new gene editing technology outside of CRISPR-Cas, you wouldn’t have to pay license fees to other companies,” he said. “I want to be one of the scientists to come up with the next technology after CRISPR. I plan to continue researching until I retire—at the very least, I hope to be able to contribute to the research community by discovering more useful enzymes.”
This article was first published in the July 2020 print version of Asian Scientist Magazine.
Click here to subscribe to Asian Scientist Magazine in print.
———
Copyright: Asian Scientist Magazine.
Disclaimer: This article does not necessarily reflect the views of AsianScientist or its staff.