AsianScientist (Nov. 17, 2016) – Using a modified version of the gene editing enzyme Cas9, researchers at the Institute of Microbiology of the Chinese Academy of Sciences (IMCAS) have developed a universal pathogen detection system. Their work has been published in ACS Synthetic Biology.
Fast and reliable detection of specific bacterial or viral nucleic acid sequences is urgently required in health care as well as in food and environmental safety monitoring. Current in vitro nuclear acid detection methods mainly rely on polymerase chain reaction-based technology. Developing specific primers or reporter systems can take a long time, requires costly equipment and may not be accurate due to non-specific amplification.
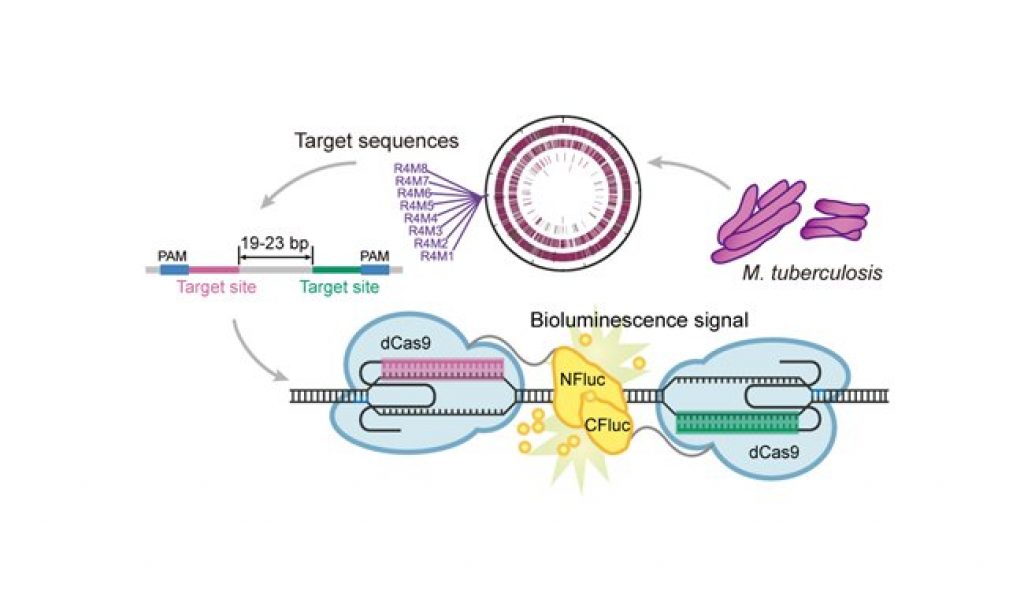
Instead, a group of researchers led by Professor Lou Chunbo fused split subunits of the luciferase enzyme to a pair of dCas9 proteins to form a specific reporter system. This reporter system is programmed by single guide RNAs complementary to the upstream and downstream segments of a target DNA sequence. When substrate DNA containing the two segment-sequences in proximity are detected and bound by the pair of dCas9 proteins, luminescence is generated from the catalytic activity of the joined luciferase subunits.
As a proof of concept, the researchers successfully distinguished Mycobacterium tuberculosis genome markers. They then selected a number of marker sequences from the large pool for high-throughput screening and succeeded in specifically detecting pathogen DNA, demonstrating the feasibility of their in vitro pathogen detection system.
The researchers are now optimizing and upgrading the paired-dCas9 system to enrich its output signal pattern, improve detection efficacy and simplify the detection equipment and operating environment.
The article can be found at: Zhang et al. (2016) Paired Design of dCas9 as a Systematic Platform for the Detection of Featured Nucleic Acid Sequences in Pathogenic Strains.
———
Source: Chinese Academy of Sciences.
Disclaimer: This article does not necessarily reflect the views of AsianScientist or its staff.












