AsianScientist (Jun. 9, 2015) – A group from the Center for RNA Research of the Institute for Basic Science (IBS) and School of Biological Sciences in Seoul National University (SNU) has detailed how the RNA microprocessor, the DROSHA-DGCR8 complex, precisely determines cleavage sites on microRNA-containing primary transcripts, allowing faithful initiation of microRNA biogenesis.
The group’s findings, published in Cell, not only reveal the function of each part of human microprocessor, but also outline future work on the molecular structure of the protein complex which will enable various new applications of RNA interference technology in basic research and human therapeutics.
MicroRNAs are short RNA species of ~22 nucleotides that play critical roles in a wide variety of cellular processes including stem cell differentiation and tumorigenesis. The specificity of microRNA gene silencing ability is dependent on the sequence, which in turn is acquired through the miRNA production process calledmiRNA biogenesis.
MicroRNA biogenesis is initiated in the nucleus by the microprocessor, a complex of the catalytic subunit DROSHA and a co-factor DGCR8, which cleaves primary transcripts containing microRNA sequences (pri-miRNA). It is thought that the pri-miRNA processing step determines the sequences of miRNAs and thereby their actions.
The researchers from IBS and SNU, led by Dr. Woo Jae-Sung and Dr. V. Narry Kim, have made a significant advancement towards understanding the molecular mechanism of miRNA processing by using the highly pure recombinant microprocessor, which was unavailable until now.
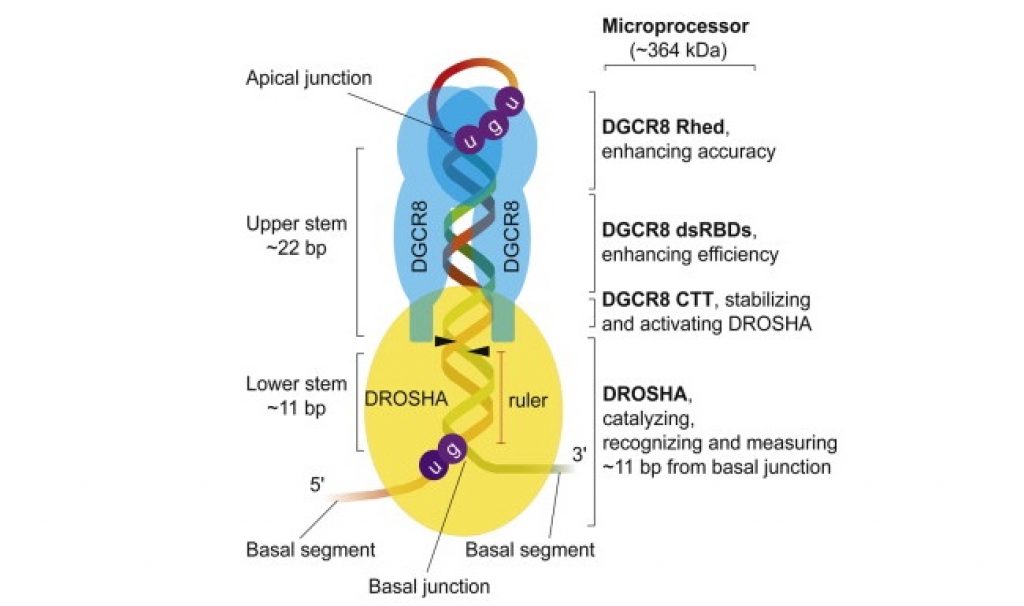
They discovered that the microprocessor consists of one DROSHA and two DGCR8 molecules. They also disclosed an important and surprising role of DROSHA as a “molecular ruler” by showing that DROSHA can recognize the ssRNA-dsRNA junction at the lower side of pri-miRNA and measure the distance of ~11 base pairs from the junction to find the precise cleavage sites.
DROSHA was also found to specifically recognize the UG motif located at the lower junction, allowing it to interact with pri-miRNAs more specifically. According to previous research, DGCR8 was found to have three functionally distinct parts: the tail to stabilize DROSHA, the body to enhance the processing efficiency by recruiting pri-miRNA and the head to ensure the processing accuracy by recognizing the upper elements of pri-miRNA including the apical UGU motif.
Based on the biochemical, biophysical and bioinformatical data presented in the study, the researchers proposed a model describing how the functional parts of the microprocessor interact with the cis-acting elements on pri-miRNA for accurate processing. The model explains how the microprocessor acts differently on various pri-miRNA substrates with different sequence and structural features and clarifies decade-standing controversies over the pri-miRNA processing mechanism.
The article can be found at: Nguyen et al. (2015) Functional Anatomy Of The Human Microprocessor.
———
Source: Institute for Basic Science.
Disclaimer: This article does not necessarily reflect the views of AsianScientist or its staff.