AsianScientist (Apr. 17, 2015) – Researchers have identified two RNA sequences that are crucial regulators of trans-splicing in flies. Their research has been published in Genes & Development.
RNA splicing is one of the essential steps of RNA processing during eukaryotic gene expression and regulation. Unlike the typical cis-splicing, trans-splicing joins exons from two separate transcripts to produce chimeric mRNA and has been detected in most eukaryotes. Trans-splicing in trypanosomes and nematodes has been characterized as a spliced leader RNA-facilitated reaction. However, its mechanism in higher eukaryotes remains unclear.
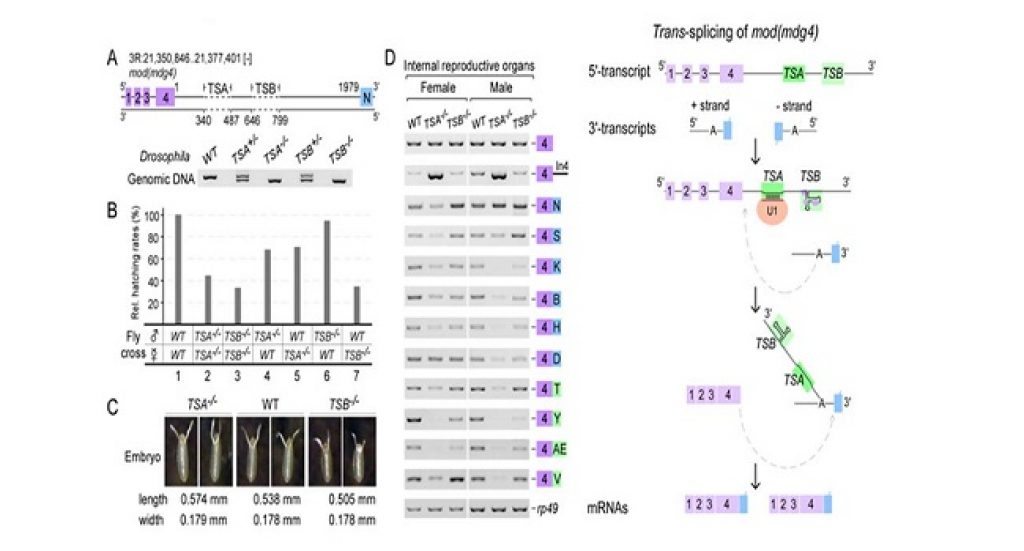
In a recent study, researchers led by Dr. Xu Yong-Zhen at the Institute of Plant Physiology and Ecology, Shanghai Institutes for Biological Sciences, Chinese Academy of Sciences, found two critical intronic RNA sequences, TSA and TSB, that promote the trans-splicing of mod (mdg4), a classic trans-spliced gene in Drosophila.
TSA, a 13-nucleotide core motif conserved across Drosophila species, is essential and sufficient for trans-splicing. TSB forms a conserved secondary structure and acts as an enhancer, which binds to chromatin factors and small nuclear RNA (snRNA) components.
Precise deletions of TSA and TSB using the CRISPR/Cas9 system resulted in developmental and viability defects in flies. By compensating TSA mutants with U1 snRNA, the researchers were able to rescue the mutant phenotype, demonstrating that U1 recruitment is critical to promote trans-splicing in vivo. Furthermore, TSA core-like motifs are also found in many other trans-spliced genes in Drosophila.
These findings reveal a general mechanism of trans-splicing in higher eukaryotes whereby the intronic RNA motifs recruit spliceosomal factors and bring separate transcripts into close proximity to promote trans-splicing.
The article can be found at: Gao et al. (2015) A Conserved Intronic U1 snRNP-binding Sequence Promotes trans-splicing in Drosophila.
———
Source: Shanghai Institutes for Biological Sciences.
Disclaimer: This article does not necessarily reflect the views of AsianScientist or its staff.